Cross-Kingdom Conservation of Sleep-Related Genes in Chlamydomonas Sheds Light on Fundamental Biological Process
Mysuru, Karnataka, 3 April 2025 — A groundbreaking study published in the Journal of Cellular Biochemistry has uncovered compelling evidence that sleep, a behavior once thought to be exclusive to complex organisms, may have its roots in the simplest forms of life. The research, led by an international team of scientists, identifies 112 putative sleep-related genes in the single-celled green alga Chlamydomonas reinhardtii, suggesting that the molecular mechanisms regulating sleep have been conserved across kingdoms for over 500 million years.
Key Findings:
- Evolutionary Conservation: The study reveals that sleep-related genes in humans share significant similarities with those in Chlamydomonas, highlighting an evolutionary continuum from unicellular organisms to mammals.
- Novel Discoveries: Among the identified genes are previously uncharacterized proteins, such as CHLRE_16g686650v5, which exhibit high sequence similarity to human sleep genes, opening new avenues for understanding sleep regulation.
- Functional Insights: Well-known proteins like casein kinase I (CKI) and voltage-dependent calcium channels, critical for circadian rhythms and signaling in humans, were also found in Chlamydomonas, reinforcing their pleiotropic functions.
Implications for Science and Medicine:
This research challenges traditional views of sleep as a purely neural phenomenon, proposing instead that its origins lie in fundamental cellular processes. “Our findings suggest that sleep’s essential functions—energy conservation, cellular repair, and adaptation—may have evolved early in life’s history,” said Pandi-Perumal, one of the study’s lead authors. “Our current findings related to putative sleep genes in Chlamydomonas, a flagellated autotrophic plankton, could revolutionize how we view the evolution of human sleep genes, which eventually helps to study sleep disorders and develop potential therapies by targeting conserved pathways.”
Why It Matters:
- Model Organism: Chlamydomonas, a simple alga with a well-mapped genome, offers a powerful tool for dissecting sleep mechanisms without the complexity of animal models.
- Future Research: The study paves the way for exploring how these putative sleep genes in single-celled organisms contribute to survival, with potential applications in neurology, chronobiology, and even synthetic biology.
Expert Commentary:
“If sleep orthologs exist in algae, it forces us to rethink its purpose,” noted Dr. Chidambaram, co-corresponding author and a professor at the Department of Pharmacology, JSS College of Pharmacy, JSS Academy of Higher Education and Research, Mysuru, Karnataka, India. “This work bridges the gap between microbiology and neuroscience, offering a unified framework to decode sleep’s mysteries through phylogenomics.”
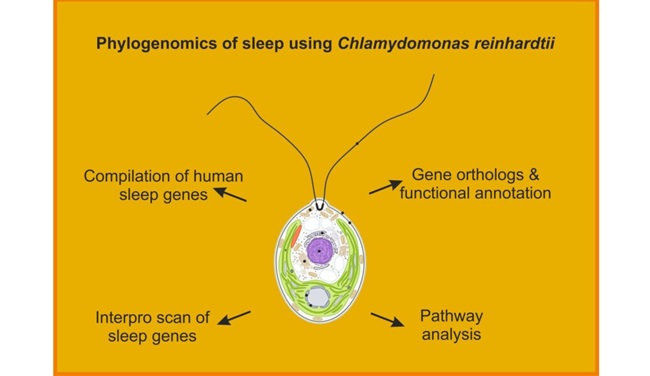
According to Dr. Saravanan, one of the co-authors and Bioinformatics expert, “This study presents an interesting computational framework that enhances cross-genomics comparison by integrating multiple phylogenomic analysis of sleep genes such as Gene ontology, Interpro sites analysis, Repeats analysis and Pathway analysis, offering significant improvements in accuracy, efficiency, and analytical depth.”
According to Prof. Abraham, a Former Professor & Head of the Post-Graduate Department of Botany, at The American College, Madurai, who is one of the co-authors of the study said, “This is an exemplary work, even though we say it ourselves. This phylogenomics study unveils unprecedented insights into evolutionary and functional parallels between putative sleep genes of Chlamydomonas and human sleep genes.”
Overall, the results of this study present an interesting and in-depth analysis of Cross-Kingdom Genomic Conservation of Putative Human Sleep-Related Genes in Chlamydomonas, uncovering previously unreported evolutionary and functional parallels, particularly in protein kinase domains involved in the information storage and processing, providing new insights into conserved genetic elements and their implications in cellular processes.
Publication Details:
The full study, “Cross-Kingdom Genomic Conservation of Putative Human Sleep-Related Genes: Phylogenomic Evidence From Chlamydomonas reinhardtii,” is available in the Journal of Cellular Biochemistry (2025; DOI: [https://doi.org/10.1002/jcb.70030]).
For Media Inquiries:
Contact: Lemuria Global Network
Email: lemuriaglobal@gmail.com
About the Journal of Cellular Biochemistry:
A peer-reviewed publication by Wiley, the journal advances research on cellular mechanisms underpinning health and disease. https://onlinelibrary.wiley.com/doi/full/10.1002/jcb.70030
How to cite this paper:
Pandi-Perumal SR, Saravanan KM, Paul S, Abraham GC, Warren Spence D, Chidambaram SB. Cross-Kingdom Genomic Conservation of Human Sleep-Related Gene Orthologs: Phylogenomic Evidence from Chlamydomonas reinhardtii. J Cellular Biochem. https://doi.org/10.1002/jcb.70030 (First published: 31 March 2025).